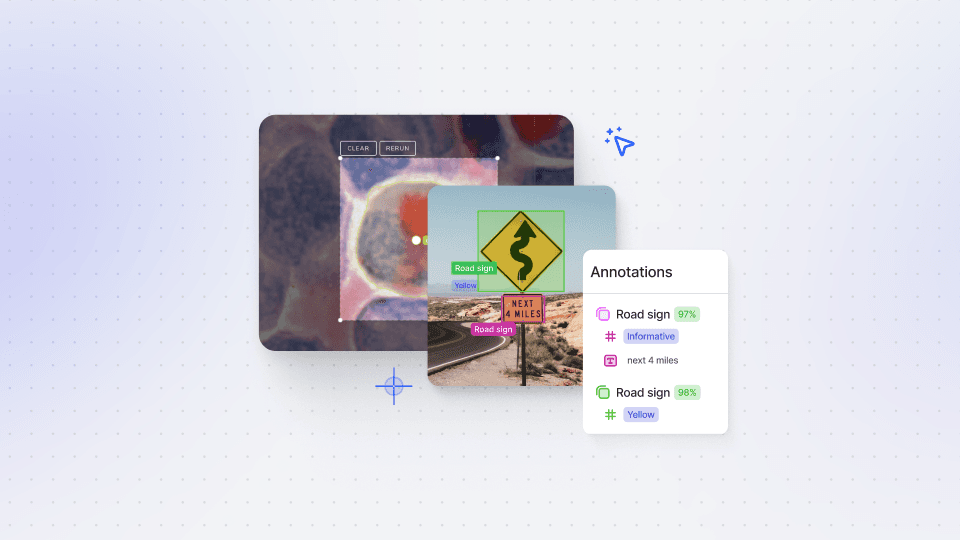
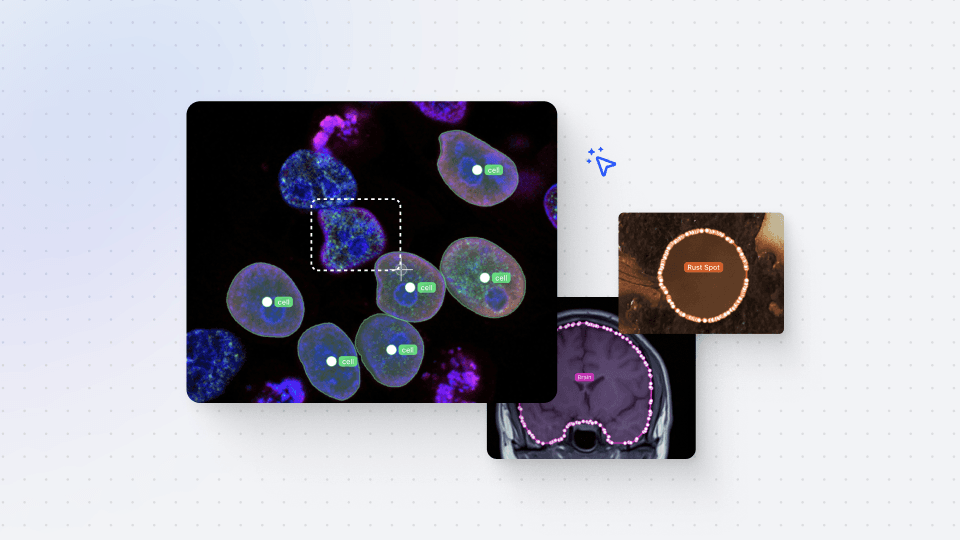
Annotations Import
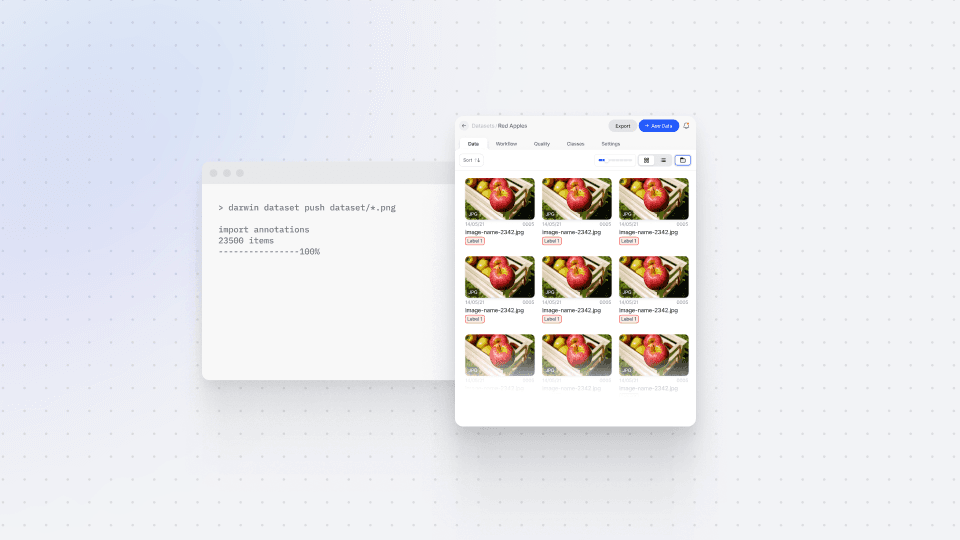
Play video
7:22
In this Darwin Advanced session, we discuss how you can import annotations to V7 by using the command line interface (CLI) or the software development kit (SDK) - in any of our supported file formats. Head to our Import Annotations documentation to access a detailed overview of the process and the scripts discussed in the video.
This tutorial begins with a breakdown of the Darwin JSON file format, the most versatile and widely used option for importing annotations. However, if you require another format - don’t worry, we support COCO, YOLO, Instances PNG, CVAT, and many others.
We begin by breaking down the makeup of the Darwin JSON format, highlighting the annotation and item data specific to the image or video you’re importing. To get started with annotations import, we showcase two methods: the CLI and the SDK.
The step-by-step run-through of the CLI method includes how users can authenticate themselves using the “darwin authenticate” command, and how they can access datasets by using “darwin datasets remote”.
Next, we demonstrate the process of using the SDK, with code examples. Here we walk you through initiating the client and specifying the annotation format, to uploading the annotations to the dataset hosted on V7.
You’ll come away with a detailed and accessible explanation of how to import annotations to V7, allowing you to organize and drive your computer vision projects forward.